MEEP
Meep (or MEEP) is a free finite-difference time-domain [1] simulation software package developed at MIT to model electromagnetic systems. A number of people, including @aero, have been using it to simulate the evolution of the fields within the EM Drive.
Contents
Setup
Details regarding setting up Meep for EM Drive simulations are provided below. Note: this whole section should not be needed, if the Meep folks provided clear instructions/deployment packages/deployment options. Someone could point them to this page...
Most people are running Meep on Ubuntu, although @lmbfan has also managed to set it up on Windows via Cygwin.[2]
Ubuntu Linux setup
Default installation
apt-get install meep h5utils
or, for the multi-processor version (potential performance improvement, see Runtime & Performance section, below)
apt-get install meep-mpi h5utils
The installation for a single machine is described here. Parallel/cluster installation instructions are documented here
Manual installation
@leomillert recommended[3] the following:
- Meep 1.3 (compiled & installed from its sources[4])
- libctl 3.2.2 (compiled & installed from its sources)
- Guile 2.0.11 (id)
- Harminv 1.4 (id)
- OpenBLAS 0.2.12 (installed from its binaries)
- HDF5 1.9.224 (compiled & installed from its sources)
- h5utils 1.12.1 (id)
Cloud computing: Amazon EC2 AMI
Due to the long computing times required to simulate the frustum until it has reached a steady state solution, you may find it more convenient to run the simulation on Amazon EC2 than on your machine.
An Amazon machine image (AMI) with Meep pre-installed has been made available by @dumbo; the name is ubuntu-trusty-meep-emdrive and the AMI ID is ami-e5cbc9d5. The AMI is based on Ubuntu 14.04.2 (Trusty Tahr). Meep was installed from source using the following dependencies:
- Meep 1.3 (installed from source)
- libctl 3.2.2 (installed from source)
- Guile 2.0.9 (as provided by Ubuntu package guile-2.0)
- Harminv 1.4 (installed from source)
- OpenBLAS 0.2.14 (installed from source)
- HDF5 1.8.15 Patch 1 (installed from source)
- h5utils 1.12.1 (installed from source)
- GSL 1.16 (installed from source)
Virtualization: Virtualbox image
TBD: Describe here where to find a Virtualbox pre-build image. One pre-built Meep image for VBox was provided by Meeper, Quixote who posts on NSF.
A Virtualbox image will allow running Meep using the same setup under Linux, Windows, Mac, Solaris The overhead of a Virtualbox is negligible compared to the benefits (i.e. ease of installation).
Quixote's VBox Meep install is available here, first some notes:
Note: Meep and ancillary codes were compiled from the latest source, as of July, 2015.
Note: Most important - Go to the following listed Google drive page, then read the instructions.
Note: Using a virtual machine running Meep installed as directed, expect about 3% overhead compared to the same computer running Meep installed on bare metal.
Note: This overhead is also negligible compared to the overhead of a generic pre-compiled download package.
Note: some HP BIOSES have a glitch, When You ENABLE VT-X/AMD-V, You DISABLE it, so do try both ways if necessary.
Emulation: Bochs image
Currently, it is believed that .h5 files produced by Meep are machine-specific, and hence cannot be compared exactly. Note that this has not been demonstrated yet, and that other artifacts or side-effects may lead to differences in .h5 files.
Bochs allows emulating Intel x86 CPUs.
A Bochs image would provide an opportunity to produce exactly the same results (ie .h5 files) on different (physical) machines. This will in turn allow establishing formally that a setup is correct, trough, inter alia, comparison of the hash value of said .h5 files. It would also allow perfect reproducibility of the results.
Unfortunately, Bochs comes with a performance hit (TBD: quantify).
Docker
There exists a proposal to use Docker for deployments of Meep. The expected benefits are unclear at this stage.
Usage
The instructions below were provided by @leomillert:[5]
1. Execute
wget "https://forum.nasaspaceflight.com/index.php?action=dlattach;topic=37642.0;attach=1042821" -O NSF-1701.ctl meep NSF-1701.ctl
After completion of the execution, Meep outputs nine .h5 files (this may take a long time: see below).
2. Create CSV files
h5totxt -t 13 -0 -y -0 ex.h5 > zCopper-exy.csv
3. Open zCopper-exy.csv in a spreadsheet and aero's zCopper-exy.csv (TBD:provide location of aero csv) on another. Open a third spreadsheet that is one spreadsheet values minus the other, entry by entry. Check the highest entry (in absolute value) of the difference. If it's negligible you are good to go. If it's a value too big (greater than 10^-6), your Meep installation isn't in sync with ours, so it's of no use.
4. Now you are good to go. Make a new directory to start the tests. Copy NSF-1701.ctl there.
5. Open NSF-1701.ctl in a text editor and change a single value. For example, (set! high 10.2) means the model is 10.2 inches high. Change the 10.2 to another value and save NSF-1701.ctl with this single change. This is called sensitivity analysis. One value at a time. (set! high 10.2) was just an example, change any value of interest
meep NSF-1701.ctl h5totxt -t 13 -0 -y -0 ex.h5 > zCopper-exy.csv
6. Compare your new zCopper-exy.csv with your old one. See if there was any relevant change (do the spreadsheet comparison again). If there is no significant change in values, it means the modification made doesn't impact the behaviour of the EMDrive. This is an important information for scientists, so let us know. Otherwise, if there was a significant change, let us know if it was positive or negative and its intensity. If you don't know how, just upload the .h5 files somewhere and we will analyse it.
Validating Meep solutions
It would be interesting to simulate the behaviour of EMDrive using a cylindrical shape, for which an exact solution can be calculated. This would not only allow to validate that Meep is predicting things correctly, but also would allow to validate that the units are correct (Meep has a peculiar way of representing units...).
Candidates for simulations
Meepers may wish to investigate variations of the reference model (NSF-1701.ctl) and report on their results. The following variations are suggested:
Sensitivity analysis
These parameters would benefit from sensitivity analysis:
- Frequency (from 2.3 Ghz to 2.6 Ghz with 0.01 Ghz increments)
- Height
- Diameter of the small base
- Diameter of the large base
- Wall thickness
- Antenna location
An animation showing the change in each field (including the Poynting field) at some well chosen time t and slice s can help visualizing the results and determining if small changes in values affect the results significantly (or not).
The golden ratio has been proposed, although it is not clear to what it applies (height/small diameter, height/big diameter, big diameter/small diameter/...)
Change of outside material
- Copper
- Silver
- Aluminum
- Gold (plated!)
- Copper side, (one) iron base
- Copper side, (one) Metglas® 2714A base
Change of inside material
- Air
- Vacuum
- Sulfur hexafluoride
- Water
- Oil
- Ammonia
Change of shape
- Hull
- Parabolic
- Exponential
- Hyperbolic
- Tractrix
- Hexagonal section
- Square section
- Antenna
- Loop
- Circular
- Square
- Fractal
- Dipole
- Loop
Other type of change
- Small holes on sides only
- Small holes on large base only
- Small holes on the complete outer hull
- Curved (ie spherical end) instead of flat ends
- Increasing resolution and/or lattice
- (More) accurate modelling of current source (ie use custom-src instead of continuous / Gaussian source)
- Direct measurements of flux spectra (define transmission (add-flux ...))
- Direct measurements of force spectra (add-force fcen df nfreq force-regions...)
- Using symmetries to speed up calculations
Improving Meep output
Converting Meep csv to an image with contour and scale
The following Python script (requires MatPlotLib) converts Meep .csv output to an image with contours (16) and a vertical scale:
import numpy as np
import matplotlib.pyplot as plt
data = np.genfromtxt('13-eyz30.csv',delimiter=',',skip_header=0)
data[data == 0] = 'nan'
plt.title('Ey z - 30',fontsize=13)
frame=plt.gca()
frame.axes.set_xticklabels([])
frame.axes.set_yticklabels([])
plt.xticks(np.arange(0, len(data[0]), 10.0))
plt.yticks(np.arange(0, len(data), 10.0))
plt.grid(b=True, which='both', color='0.65', alpha=0.2, linestyle='-')
cs = plt.contourf(data, 16, cmap=plt.cm.RdYlBu, antialiased=True)
cb=plt.colorbar(cs)
cb.ax.tick_params(labelsize=10)
plt.savefig('eyz30.png',bbox_inches='tight')
plt.show()
Result:

Note: When I have a bit of time, I'll update this script to read directly from the HDF5 file, alleviating the need for the intermediary .csv. This will both be faster and ensure that data is read as close as possible from the source. I'll also create a .mp4 automatically from the HDF5 data.
Color maps
Meepers may wish to explore different color maps to represent the outputs. The Meep h5topng utility supports color maps (http://ab-initio.mit.edu/wiki/index.php/Color_tables_in_h5topng), with bluered and dkbluered, combined with the -Z option, looking most promising. Audacious meepers may consider defining their own color scale and color bar (this is explained in the link above).
Contour plots
Meep adventurers may wish to consider improving h5topng to produce images which contain:
- The numerical value on the contour boundaries and/or
- A scale alongside the image
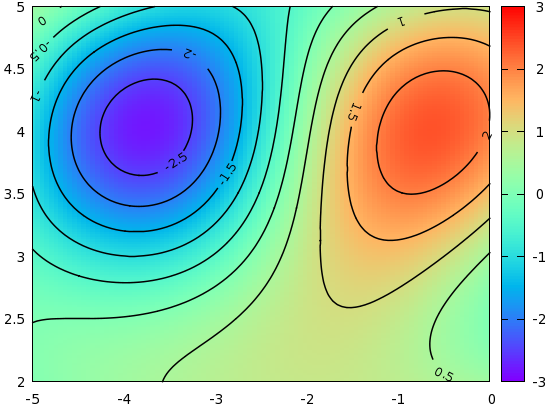
- Example 1. Contour with value and scale
- Example 2. Value on boundaries only:

- Example 3. Scale only:

- Example 4. Contour plot with boundaries on transitions:
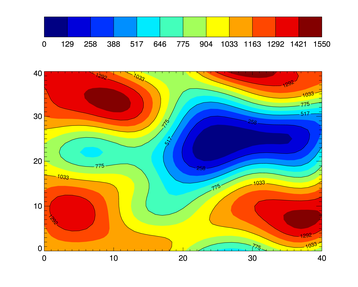
There is a Python h5topng script available here: https://github.com/NealJMD/gala-scripts/blob/master/h5topng.py. Maybe it could serve as a basis ?
Another possible venue would be use gnuplot, see here for example: http://www.phyast.pitt.edu/~zov1/gnuplot/html/contour.html Waiting for a good soul to put everything in a script...
Yet another possibility: use matplotlib to directly produce the contour plots from CSV files
- See here: http://matplotlib.org/gallery.html
- And here: http://stackoverflow.com/questions/16371725/getting-a-contour-plot-to-use-data-from-a-csv-file
Surfit
surfit (http://surfit.sourceforge.net/) is a computer program which enables to calculate regular grid from various data (scattered points, 2D and 3D contours, surfaces, etc) in different ways (interpolation, approximation, inequalities, etc). Presenting EMdrive results using it could improve presentation in a significant way. Note that surfit generates (labelled) contour plots.
Output jpg
Would an h5tojpg allow to directly produce smaller images without loosing (too much) quality?
Creating an animation from a set of images
An animation conveys more information and takes less space than individual images! Here is how to create a mp4 from a set of .pngs:
Using ImageMagick:
convert -antialias *.png emdrive.mp4
Using FFmpeg:
ffmpeg -framerate 10 -i emdrive%03d.png -s:v 1280x720 -c:v libx264 -profile:v high -crf 20 -pix_fmt yuv420p emdrive.mp4
Models & results
Repository
@tidux has set up a Git repository at http://git.emdrive.science/ with anonymous read access. [6]. PM @tidux with your SSH key if you want write access.
Models for consideration
- deuteragenie toy 2D model (TBD find ctl, movie on thread 2)
- aero bradycone 3D model (TBD find refs.)
- aero nsf 3d model (TBD find refs.)
- leo? 3d model with full synchronization and updates for Meep 1.3 syntax (TBD find refs.)
- A a 2D centered-slice model would allow for fast simulations/testing before moving up to the more time consuming 3D simulations.
- Tajmar
Runtime & performance
server 'pollux': Compaq Elite 8300, Intel i5-3470 @ 3.2 Ghz, 24GB DDR3-1600, Debian 8.1, utilizing NSF-1701.ctl as a reference configuration:
- one processor, non-mpi version: 75 minutes (1:15:07)
- one processor, mpi: 72 minutes (1:12:12)
- four processors: 50 minutes (49:22)
- two simultaneous runs, each using two processors: 100 minutes (1:39:22)
TBD: add more specs, make a nice table Currently: count on more than one hour to run the reference .ctl file on a "normal" computer.